"V体育官网" Long noncoding RNAs and the genetics of cancer
- PMID: 23660942
- PMCID: V体育官网 - PMC3694235
- DOI: 10.1038/bjc.2013.233
Long noncoding RNAs and the genetics of cancer
Abstract
Cancer is a disease of aberrant gene expression. While the genetic causes of cancer have been intensively studied, it is becoming evident that a large proportion of cancer susceptibility cannot be attributed to variation in protein-coding sequences VSports手机版. This is highlighted by genome-wide association studies in cancer that reveal that more than 80% of cancer-associated SNPs occur in noncoding regions of the genome. In this review, we posit that a significant fraction of the genetic aetiology of cancer is exacted by noncoding regulatory sequences, particularly by long noncoding RNAs (lncRNAs). Recent studies indicate that several cancer risk loci are transcribed into lncRNAs and these transcripts play key roles in tumorigenesis. We discuss the epigenetic and other mechanisms through which lncRNAs function and how they contribute to each stage of cancer progression, understanding of which will be crucial for realising new opportunities in cancer diagnosis and treatment. Long noncoding RNAs play important roles in almost every aspect of cell biology from nuclear organisation and epigenetic regulation to post-transcriptional regulation and splicing, and we link these processes to the hallmarks and genetics of cancer. Finally, we highlight recent progress and future potential in the application of lncRNAs as therapeutic targets and diagnostic markers. .
Figures

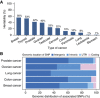
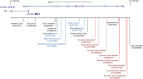
References (V体育2025版)
-
- Amaral PP, Dinger ME, Mercer TR, Mattick JS. The eukaryotic genome as an RNA machine. Science. 2008;319:1787–1789. - PubMed
-
- Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, Kodzius R, Shimokawa K, Bajic VB, Brenner SE, Batalov S, Forrest ARR, Zavolan M, Davis MJ, Wilming LG, Aidinis V, Allen JE, Ambesi-Impiombato A, Apweiler R, Aturaliya RN, Bailey TL, Bansal M, Baxter L, Beisel KW, Bersano T, Bono H, Chalk AM, Chiu KP, Choudhary V, Christoffels A, Clutterbuck DR, Crowe ML, Dalla E, Dalrymple BP, de Bono B, Della Gatta G, di Bernardo D, Down T, Engstrom P, Fagiolini M, Faulkner G, Fletcher CF, Fukushima T, Furuno M, Futaki S, Gariboldi M, Georgii-Hemming P, Gingeras TR, Gojobori T, Green RE, Gustincich S, Harbers M, Hayashi Y, Hensch TK, Hirokawa N, Hill D, Huminiecki L, Iacono M, Ikeo K, Iwama A, Ishikawa T, Jakt M, Kanapin A, Katoh M, Kawasawa Y, Kelso J, Kitamura H, Kitano H, Kollias G, Krishnan SPT, Kruger A, Kummerfeld SK, Kurochkin IV, Lareau LF, Lazarevic D, Lipovich L, Liu J, Liuni S, McWilliam S, Madan Babu M, Madera M, Marchionni L, Matsuda H, Matsuzawa S, Miki H, Mignone F, Miyake S, Morris K, Mottagui-Tabar S, Mulder N, Nakano N, Nakauchi H, Ng P, Nilsson R, Nishiguchi S, Nishikawa S, Nori F, Ohara O, Okazaki Y, Orlando V, Pang KC, Pavan WJ, Pavesi G, Pesole G, Petrovsky N, Piazza S, Reed J, Reid JF, Ring BZ, Ringwald M, Rost B, Ruan Y, Salzberg SL, Sandelin A, Schneider C, Schönbach C, Sekiguchi K, CAM Semple, Seno S, Sessa L, Sheng Y, Shibata Y, Shimada H, Shimada K, Silva D, Sinclair B, Sperling S, Stupka E, Sugiura K, Sultana R, Takenaka Y, Taki K, Tammoja K, Tan SL, Tang S, Taylor MS, Tegner J, Teichmann SA, Ueda HR, van Nimwegen E, Verardo R, Wei CL, Yagi K, Yamanishi H, Zabarovsky E, Zhu S, Zimmer A, Hide W, Bult C, Grimmond SM, Teasdale RD, Liu ET, Brusic V, Quackenbush J, Wahlestedt C, Mattick JS, Hume DA, Kai C, Sasaki D, Tomaru Y, Fukuda S, Kanamori-Katayama M, Suzuki M, Aoki J, Arakawa T, Iida J, Imamura K, Itoh M, Kato T, Kawaji H, Kawagashira N, Kawashima T, Kojima M, Kondo S, Konno H, Nakano K, Ninomiya N, Nishio T, Okada M, Plessy C, Shibata K, Shiraki T, Suzuki S, Tagami M, Waki K, Watahiki A, Okamura-Oho Y, Suzuki H, Kawai J, Hayashizaki Y, FANTOM Consortium, RIKEN Genome Exploration Research Group and Genome Science Group (Genome Network Project Core Group) The transcriptional landscape of the mammalian genome. Science. 2005;309:1559–1563. - "VSports注册入口" PubMed
-
- Chan KC, Jiang P, Zheng YW, Liao GJ, Sun H, Wong J, Siu SS, Chan WC, Chan SL, Chan AT, Lai PB, Chiu RW, Lo YM. Cancer genome scanning in plasma: detection of tumor-associated copy number aberrations, single-nucleotide variants, and tumoral heterogeneity by massively parallel sequencing. Clin Chem. 2013;59:211–224. - PubMed
Publication types
- Actions (VSports最新版本)
- V体育官网入口 - Actions
MeSH terms
- V体育官网 - Actions
- V体育2025版 - Actions
- "VSports app下载" Actions
- "V体育安卓版" Actions
- VSports手机版 - Actions
Substances
- "V体育安卓版" Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources (V体育平台登录)

